SAMENT scRNA-seq Resource
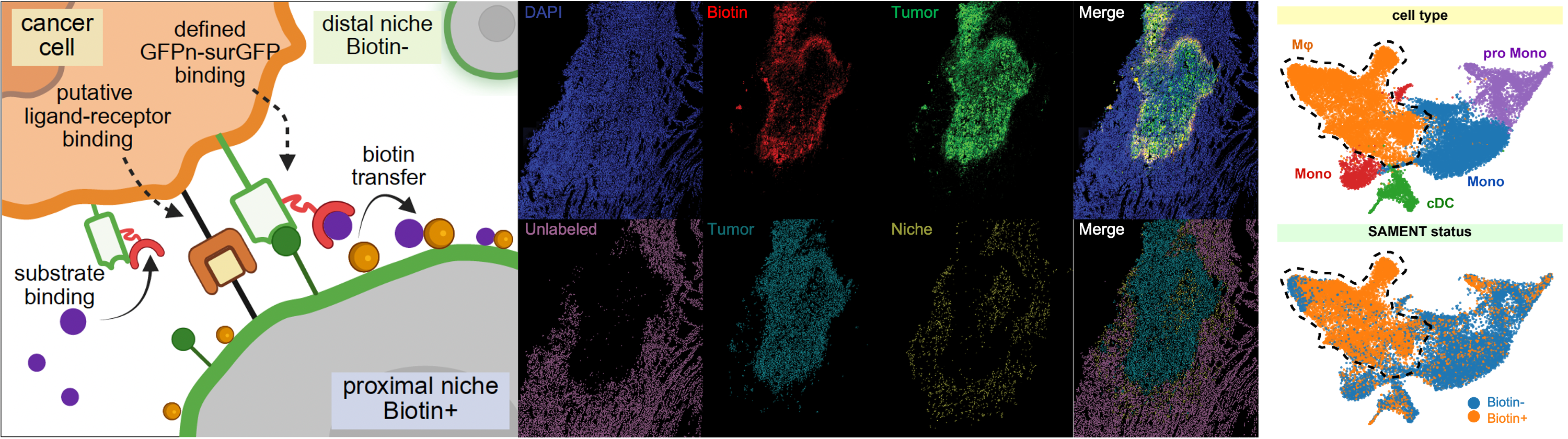
Unbiased metastatic niche-labeling identifies estrogen receptor-positive macrophages as a barrier of T cell infiltration during bone colonization
This repository provides instructions and code to reproduce the analyses, numerics, and figures associated with the SAMENT scRNA-seq study.
Interactive Exploration
Interactive volcano plots allow exploration of differentially regulated pathways between biotin-positive and biotin-negative immune populations.
Analysis Pipeline
| File | Description |
|---|---|
| 01_preprocessing_demultiplexing_template.Rmd | Process Cell Ranger outputs and generate individual Seurat objects |
| 02_integration.Rmd | Integration of the single-cell objects |
| 03_scanpy_plot.ipynb | Generate plots using Scanpy |
| 04_processing_for_DESeq2.Rmd | Prepare DESeq2 object for DEG and pathway analysis |
| 05_DESeq2_DEG.Rmd | Differential gene expression analysis |
| 06.2_GSEA.Rmd | Gene set enrichment analysis |
| 07.GSVA.Rmd | GSVA pathway activity analysis |
| 08.velocyto.ipynb | RNA velocity trajectory analysis |
| 09.expression_distence.ipynb | Transcriptomic distance analysis |
| 10.CellChat_comparsion.Rmd | Cell-cell communication analysis |
Citation
Xu Z, Liu F, Ding Y, et al. Unbiased metastatic niche-labeling identifies estrogen receptor-positive macrophages as a barrier of T cell infiltration during bone colonization. Cell (accepted).